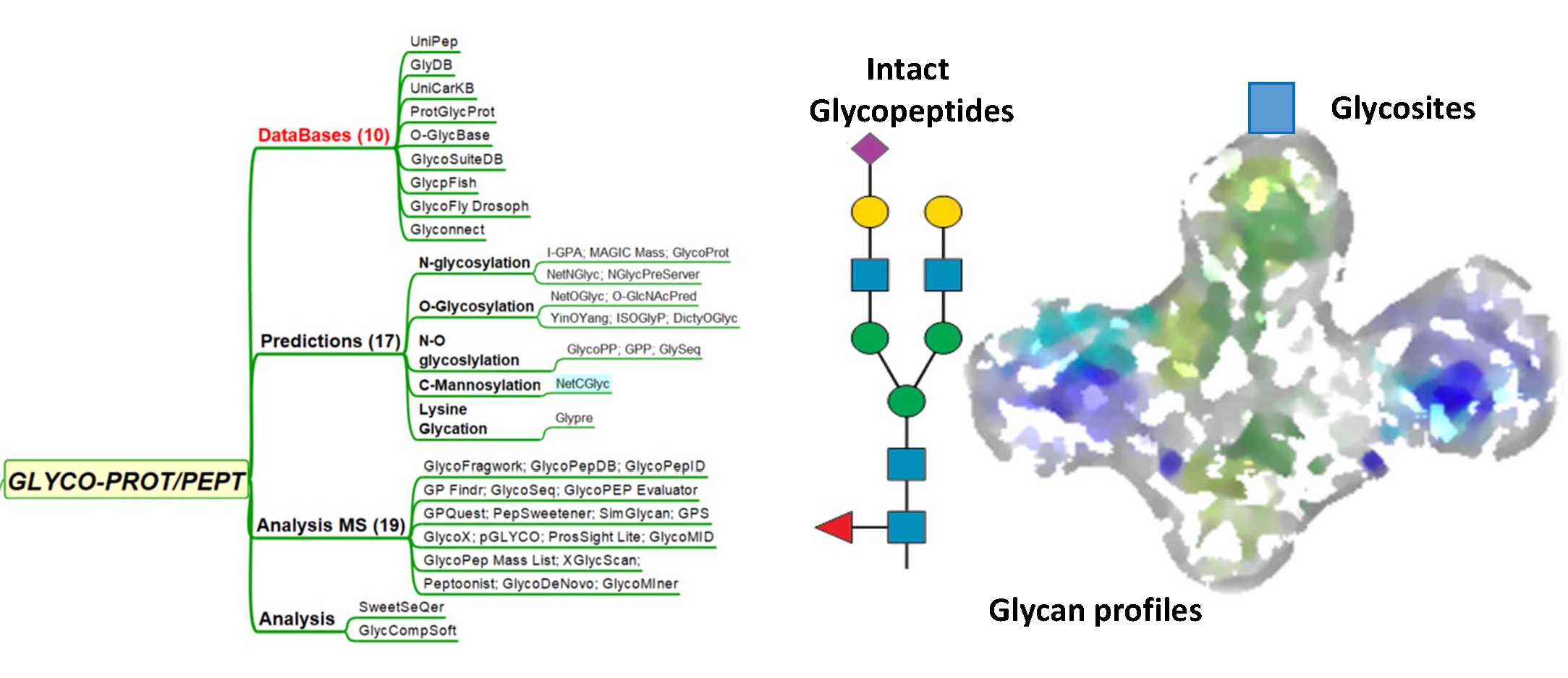
DATABASES
- Glyconnect https://glyconnect.expasy.org/ GlyConnect is a platform integrating sources of information to help characterise the molecular components of protein glycosylation
- GlyDB N-Glycan Structure Annotation of Glycopeptides Using a Linearized Glycan Structure Database
- GlycoFish http://betenbaugh.jhu.edu/GlycoFish A Database of Zebrafish N-linked Glycoproteins Identified Using SPEG Method Coupled with LC/MS
- GlycoFly Drosoph http://betenbaugh.jhu.edu/GlycoFly GlycoFly is a database for Drosophila N-linked glycoproteins identified using SPEG—MS techniques.
- GlycoPAT http://www. VirtualGlycome.org/glycopat GlycoProteomics Analysis Toolbox
- GlyProt http://www.glycosciences.de/glyprot/ GlyProt is a web-based tool that enables meaningful N-glycan conformations to be attached to all the spatially accessible potential N-glycosylation sites of a known three-dimensional (3D) protein structure.
- GlycoSuiteDB http://www.glycosuite.com GlycoSuiteDB is a relational database that curates information from the scientific literature on glyco-protein derived glycan structures
- O-GlycBase http://www.cbs.dtu.dk/databases/OGLYCBASE/ A database of glycoproteins with O-linked glycosylation sites
- UniCarbKB http://unicarbkb.org UniCarbKB : New database features for integrating glycan structure abundance, compositional glycoproteomics data, and disease associations
- UniPep http://www.unipep.org A database for human N-linked glycosites : a resource for biomarker discovery
PREDICTIONS
N-Glycosylation
- I-GPA Integrated GlycoProteome Analyzer (I-GPA) for Automated Identification and Quantitation of Site-Specific N-Glycosylation.
- MAGIC http://ms.iis.sinica.edu.tw/MAGIC-web/index.html. Identifies intact N-glycosylated peptides from a public protein database without requiring any prior information of proteins or glycans
- NetNGlyc http://www.cbs.dtu.dk/services/NtNGlyc/ The NetNglyc server predicts N-Glycosylation sites in human proteins using artificial neural networks that examine the sequence context of Asn-Xaa-Ser/Thr sequons
- GlycoProt
O-Glycosylation
- DictyOGlyc http://www.cbs.dtu.dk/services/DictyOGlyc/ The DictyOGlyc server produces neural network predictions for GlcNAc. O-glycosylation sites in Dictyostelium discoideum proteins.
- ISOGlyP isoglyp.utep.edu/ Provides isoform specific O-glycosylation prediction
- NetOGlyc www.cbs.dtu.dk/services/NetOGlyc/ The NetOglyc server produces neural network predictions of mucin type GalNAc O-glycosylation sites in mammalian proteins.
- O-GlcNAcPred http://121.42.167.206/OGlcPred/ A sensitive predictor to capture protein O-GlcNAcylation sites
- YinOYang www.cbs.dtu.dk/services/YinOYang/ The YinOYang WWW server produces neural network predictions for O-ß-GlcNAc attachment sites in eukaryotic protein sequences.
N-O-Glycosylation
- GlycoPP http://crdd.osdd.net/raghava/glycopp/ GlycoPP is a webserver for predicting potential N-and O-glycosites in prokaryotic protein sequence(s)
- GPP http://comp.chem.nottingham.ac.uk/glyco/ Algorithmically prediction of N-linked and O-linked glycosylation.
- GlySeq http://www.glycosciences.de/tools/glyseq/Uses the PDB and SwissProt to perform statistical analysis of glycosylation site
C-Mannosylation
- NetCGlyc http://www.cbs.dtu.dk/services/NetCGlyc/ Prediction of mammalian C-mannosylation sites
Lysine Glycation
- Glypre http://www.cds.dtu.dk/databases/GlycateBase-1.0/ In Silico Prediction of Protein Glycation Sites by Fusing Multiple Features and Support Vector Machine
ANALYSIS
- GlycCompSoft http://www.heparin.rpi.edu Software for Automated Comparison of Low Molecular Weight Heparins Using Top-Down LC/MS Data
- SweetSeQer http://software.steenlab.org Simple de novo filtering and annotation of glycoconjugate mass spectra
ANALYSIS MS
- GlycoDENovo https://github.com/hongpengyu/GlycoDeNovoGlycoDeNovo provides an interpretation-graph which designates how to interpret each peak until the candidate topologies of precursor ion.
- GlycoFragwork http://darwin.informatics.indiana.edu/col/ A computational framework for identification of intact glycopeptides in complex samples
- GP Finder availabe from : cblebrilla@ucdavis.edu
- GlycoMiner http://www.szki.ttk.mta.hu/ms/glycominer/ A software tool to automatically identify tandem (MS/MS) spectra obtained in liquid chromatography/mass spectrometry
- GlycoMID http://proteomics.informatics.iupui.edu/software/glycomid/ A graph-based spectral alignment algorithm that can identify glycopeptides with multiple hydroxylysine O-glycosylation sites by tandem mass spectra.
- GlycoPepDB http://hexose.chem.ku.edu/sugar.php A Tool for Assigning Mass Spectrometry Data of N-Linked Glycopeptides on the Basis of Their Electron Transfer Dissociation Spectra
- GlycoPEP Detector http://glycopro.chem.ku.edu/ZZKHome.php. A Tool for Assigning Mass Spectrometry Data of N-Linked Glycopeptides on the Basis of Their Electron Transfer Dissociation Spectra
- GlycoPEP Evaluator https://desairegroup.ku.edu/research. GlycoPep Evaluator, generates decoy glycopeptides de novo and enables accurate false discovery rate analysis for small data sets
- GlycoPepID http://www.hexose.chem.ku.edu/sugar.php
- GlycoPep Mass List MassList is an application that permits to compute theoretical glycopeptide masses while creating inclusion lists for targeted data acquisition on .
- GPQuest https://www.biomarkercenter.org/gpquest A Spectral Library Matching Algorithm for Site-Specific Assignment of Tandem Mass Spectra to Intact N-glycopeptides
- GPShttp://edwardslab.bmcb.georgetown.edu/software/GlycoPeptideSearch.htmlTo explore site-specific N-glycosylation microheterogeneity of haptoglobin using glycopeptide CID tandem mass spectra and glycan database search.
- GlycoSeq https://github.com/dbaileychess/ GlycoSeq uses a heuristic iterated glycan sequencing algorithm that incorporates prior knowledge of the N-linked glycan synthetic pathway to achieve rapid glycan sequencing
- GlycoXTo Determine Simultaneously the Glycosylation Sites and Oligosaccharide heterogeneity of Glycoproteins
- pGLYCO http://pfind.ict.ac.cn/software/pGlyco1505/ pGlyco is a software tool designed for the analysis of intact glycopeptides by using mass spectrometry.
- Peptoonist Peptoonist uses tandem mass spectrometry (MS/MS) to detect glycosylated peptides and single-MS to find the N-glycans present on each of these peptides
- PepSweetener EXPASY A Web‐Based tool to support manual annotation of intact glycopeptide MS spectra
- ProSIGHT Lite http://www.prosightlite.northwestern.edu ProSight Lite : graphical software to analyze top‐down mass spectrometry data
- SimGlycan http://www.premierbiosoft.com/glycan A predictive carbohydrate analysis tool for MS/MS data